I want to save a mcmc() output so I do not need to rerun it next time. However, using saveRDS() (or save()) seems to decouple the greta_mcmc_list object from other greta_arrays.
Example code:
First, run this:
library(greta)
alpha <- normal(0, 1)
beta <- normal(0, 1)
sigma <- lognormal(1, 0.1)
y <- as_data(iris$Petal.Width)
mu <- alpha + iris$Petal.Length * beta
distribution(y) <- normal(mu, sigma)
m <- model(alpha, beta, sigma)
draws <- mcmc(m, n_samples = 500)
# save draws as rds so do not need to rerun mcmc every time
saveRDS(draws, file = "draws.rds")
Then restart the session and load the greta_mcmc_list object. This will fail halfway:
library(greta)
draws <- readRDS(file = "draws.rds")
# the codes below breaks
petal_length_plot <- seq(min(iris$Petal.Length),
max(iris$Petal.Length),
length.out = 100)
mu_plot <- alpha + petal_length_plot * beta
mu_plot_draws <- calculate(mu_plot, values = draws)
mu_est <- colMeans(mu_plot_draws[[1]])
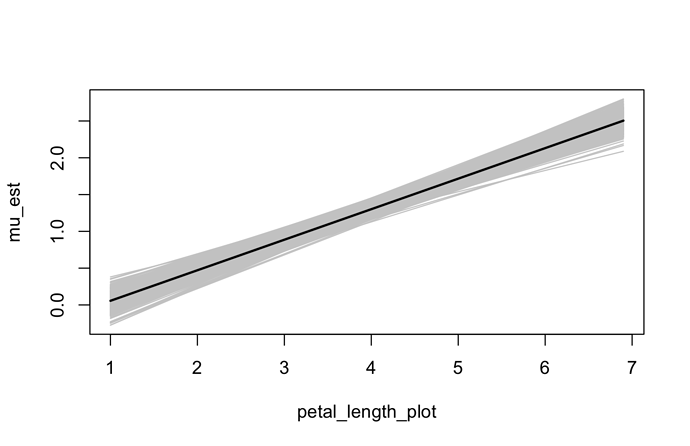
plot(mu_est ~ petal_length_plot, type = "n",
ylim = range(mu_plot_draws[[1]]))
apply(mu_plot_draws[[1]], 1, lines,
x = petal_length_plot, col = grey(0.8))
lines(mu_est ~ petal_length_plot, lwd = 2)
You should get an error:
Error: the target greta arrays do not appear to be connected to those in the greta_mcmc_list object
Did I do something wrong?